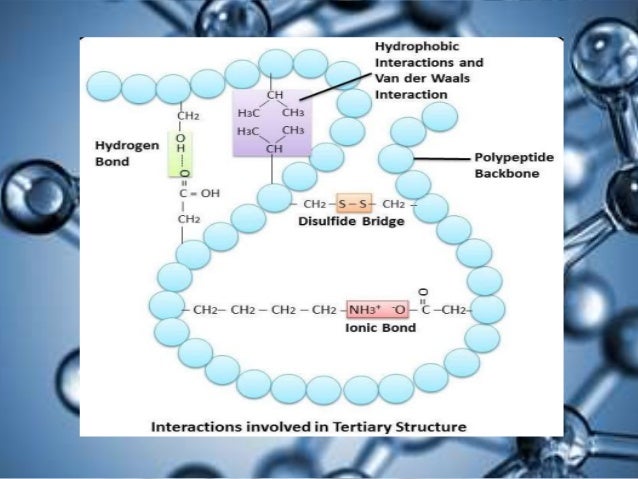
The complete list of 15N and 1H assignments is given for all the amide NH and C αH resonances of all non-proline residues, as well as the 1H assignments for some of the amino acid side chains. R represents a long complex oligosaccharide chain, joined either to ceramide where the substances are glycosphingolipids, or to the polypeptide backbone of a protein via a serine or threonine residue where the substances are glycoproteins. the amino acid sequence of its polypeptide chain secondary structure is the local spatial arrangement of a polypeptides backbone (main chain) atoms. Such a hydrogen bond is formed exactly every 4 amino. This coil is held together by hydrogen bonds between the oxygen of CO on top coil and the hydrogen of N-H on the bottom coil.  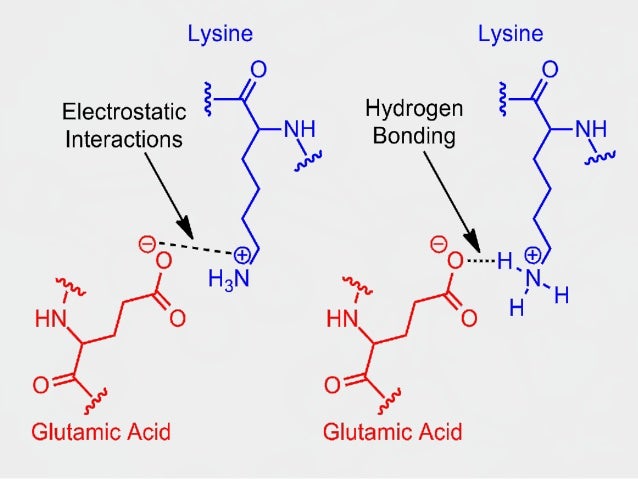
Note that the polypeptide backbone of PNA is not identical to that of natural proteins (Fig. PNA is just what its name indicates, consisting of a polypeptide backbone with nucleic acid bases attached as side chains. (The backbone just refers to the polypeptide chain apart from the R groups so all we mean here is that secondary structure does not involve R group atoms.). from publication: Encapsulation of active ingredients in polysaccharide-protein. An -helix is a right-handed coil of amino-acid residues on a polypeptide chain, typically ranging between 4 and 40 residues. Peptide nucleic acid (PNA) is a totally artificial molecule that is used as a DNA analog in genetic engineering. The limits of a residue are indicated by dashed lines, and recommended. A doubling of some peaks in the spectrum was found to be due to N-terminal heterogeneity of the 15N-labeled protein, corresponding to a mixture of wild-type and des-Ala-1-interleukin 1β. Diagrammatic representation of the structures of the H, A,and B blood group substances. Download scientific diagram Schematic of a generic polypeptide backbone. Perspective drawing of a section of polypeptide chain representing two peptide units. The authors show that the problems of amide NH and C αH chemical shift degeneracy that are prevalent for proteins of the size are readily overcome by using the 3D heteronuclear NMR technique. The fingerprint region of the spectrum was analyzed by using a combination of 3D heteronuclear 1H Hartmann-Hahn 15N- 1H multiple quantum coherence (3D HOHAHA-HMQC) and 3D heteronuclear 1H nuclear Overhauser 15N- 1H multiple quantum coherence (3D NOESY-HMQC) spectroscopies. The complete sequence-specific assignment of the 15N and 1H backbone resonances of the NMR spectrum of recombinant human interleukin 1β has been obtained by using primarily 15N- 1H heteronuclear three-dimensional (3D) NMR techniques in combination with 15N- 1H heteronuclear and 1H homonuclear two-dimensional NMR.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
